🔗 Reference
🔎 The full package documentation is available here: https://mathilde-robin.github.io/EDITH/.
🌱 The initial development of this package was motivated by the work of Tosi et al., BMC Cancer, 2018.
🎯 Aim
Characterizing interactions between drugs is an area of major interest for drug development, as exploiting synergism between drugs could allow increasing treatment efficacy using lower doses of single drugs, and avoiding antagonistic interaction is needed to maintain the therapeutic efficacy of drugs. Broadly speaking, synergism and antagonism could be defined as an increase or a reduction of the effect of a drug combination compared to the effect expected for the combination on the basis of the single agent effects.
The EDITH package enables the synergistic effects of two or three drugs to be explored, based on cell viability data obtained from combination experiments. The package is designed to analyze and visualize drug interactions and implements a widely used method for assessing drug interactions: the Bliss independence model.
⚙️ Installation
if (!requireNamespace("remotes", quietly = TRUE)) {
install.packages("remotes")
}
if (!requireNamespace("ComplexHeatmap", quietly = TRUE)) {
if (!requireNamespace("BiocManager", quietly = TRUE)) {
install.packages("BiocManager")
}
BiocManager::install("ComplexHeatmap")
}
remotes::install_github("mathilde-robin/EDITH")🔎 More details in the installation vignette: https://mathilde-robin.github.io/EDITH/articles/install.html.
📊 Input for EDITH
EDITH can explore the synergistic effects of two or three drugs based on cell viability data obtained from combination experiments. The input data should be provided in a specific format, as described in the package documentation. Example datasets are included in the package to illustrate the required format and can be found with this command:
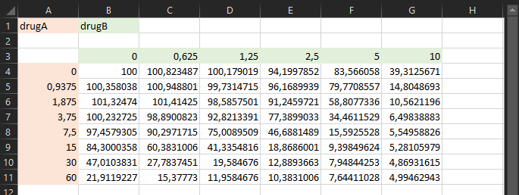
list.files(path = system.file("extdata", package = "EDITH"), full.names = TRUE)Here a snapshot of the input data format (in a excel file) for two drugs:
🔎 More details in the vignettes:
- for two drugs: https://mathilde-robin.github.io/EDITH/articles/two_drugs.html
- for three drugs: https://mathilde-robin.github.io/EDITH/articles/three_drugs.html
🚀 Usage
run_EDITH() is a all-in-one function that performs the entire analysis pipeline, from data preprocessing to visualization of results. The run_EDITH() function detects whether the input data corresponds to a two-drug or three-drug combination and applies the appropriate analysis methods accordingly. The input file path is specified via a pop-up window when the function is executed.
🎉 Output
The main outputs of the EDITH package include:
- Heatmaps visualizing the viability data across different drug concentration combinations,
- Heatmaps visualizing the expected viability based on the Bliss independence model,
- Heatmaps visualizing the interaction effect,
- Summary tables detailing the additive, combination and efficacy indexes.
Here are example heatmaps generated by EDITH for two-drug combinations: